In recent years, a number of powerful image analysis tools have been developed, including image analysis environments enabling construction of custom algorithms, such as CellProfiler 2, and more targeted software, such as tools for analysis of morphogenesis and cell shape in situ 3, 4, 5 and machine-learning enabled segmentation of twin-spot clones 6. Extracting this information, however, is time consuming and complicated. The resulting images, typically acquired by 3D confocal microscopy, are information rich and provide insights on phenotypes, such as signal intensity of reporters, clone numbers, clone size, shape and apoptosis. While this technique presents unique avenues for investigating tissue biology, extracting information from mosaic tissues by microscopic analyses is often a bottleneck. These tools come in many forms but achieve a common goal: a subset of cells within a tissue acquire a genetic alteration absent in the surrounding tissue, creating genetically distinct territories of cells. With the advent of safe, non-invasive tools for generating and marking genetically distinct subpopulations of cells within intact organisms, clonal analysis has become a common tool for the study of heterogeneous cell populations and mosaic tissues (Reviewed in 1). Furthermore, using PECAn, we identify several genes with a role in cell competition by conducting an RNAi-based screen. We find an unappreciated sexual dimorphism in Minute cell growth in competing wing discs and identify, by statistical regression analysis, tissue parameters that model and correlate with competitive death. We demonstrate the power of this pipeline by applying it to the study of Minute cell competition. 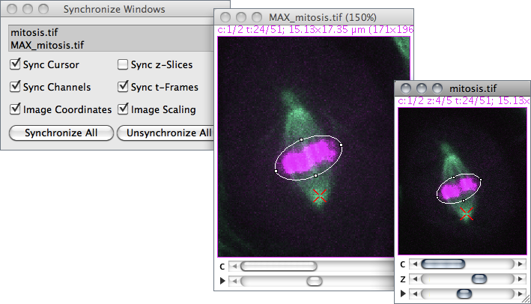
This enables researchers to perform rigorous analyses rapidly and at scale, without requiring programming skills. PECAn includes data handling, machine-learning-enabled segmentation, multivariant statistical analysis, and graph generation. Here we develop PECAn, a pipeline for image processing and statistical data analysis of complex multi-genotype 3D images. However, the 3D nature of clones makes sample image analysis challenging and slow, limiting the amount of information that can be extracted manually. Is there an easy way to do this, or am I doing something wrong?Īnother option is, I am planning to record this in a macro, since I do know the Total Wafer Area, I assume I could do some math to divide the Total void Area by the known wafer Area to get the Wafer % Voiding number I'm looking for outputted through the macro, but would need some help in how to do that.Īny help with this would be greatly appreciated.Investigating organ biology often requires methodologies to induce genetically distinct clones within a living tissue. I would like it to tell me the % of Voiding in the wafer, which would be Total Void Area\ROI Area(wafer area). But the %Area column # is low, I think because it's using the Total Area\the entire Image area. I then run the Analyze Particles routine, it surrounds the voids well and in the summary table, the Total Area column seems to come out to the correct # for Total Void Area. I then do a Threshold Color, to highlight only the voids. 
I have an image of a 4" diameter wafer with voiding. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
